This is a post in a series of tips on ontology development, see the parent post for more details.
(Note there is an excellent introduction to this topic in the ontogenesis blog)
The 2003 paper Modularisation of Domain Ontologies Implemented in Description Logics and related formalisms including OWL by Alan Rector lays out in very clear terms a simple methodology for building and maintaining compositional ontologies in a modular and maintainable fashion. From the introduction, the paper “concentrates specifically on the engineering issues of robust modular implementation in logic based formalisms such as OWL”.
Anyone involved with the authoring of ontologies should read this paper, and should strive to build modular, normalized ontologies from the outset.
The motivation for the paper is the observation that when ontologies grow beyond a certain size become increasingly hard to maintain, because polyhierarchies (i.e hierarchies where classes can have more than one parent) become increasingly “tangled”, leading to errors and high maintenance cost. This observation was based on medical ontologies such as GALEN and SNOMED, but at the time the paper came out, this was already true for many OBO ontologies such as the GO, as well as various phenotype and trait ontologies. One property all these ontologies share is their ‘compositional nature’, where more complex concepts are built up from more basic ones.

Figure: Example of difficult-to-maintain, tangled polyhierarchy, taken from the Drosophila anatomy ontology. Figure taken from OBO to OWL slides by David Osumi-Sutherland.
The methodology for “untangling” these is to decompose the domain ontology into simpler (“primitive”) ontologies, which can then be recombined using logical definitions and other axioms, and to infer the polyhierarchy using reasoning. Note that for end-users the is-a structure of the ontology remains the same. However, for the ontology maintainers, maintenance cost is much lower. This is illustrated in the paper which demonstrates the methodology using an example chemical entity hierarchy, see figure below:
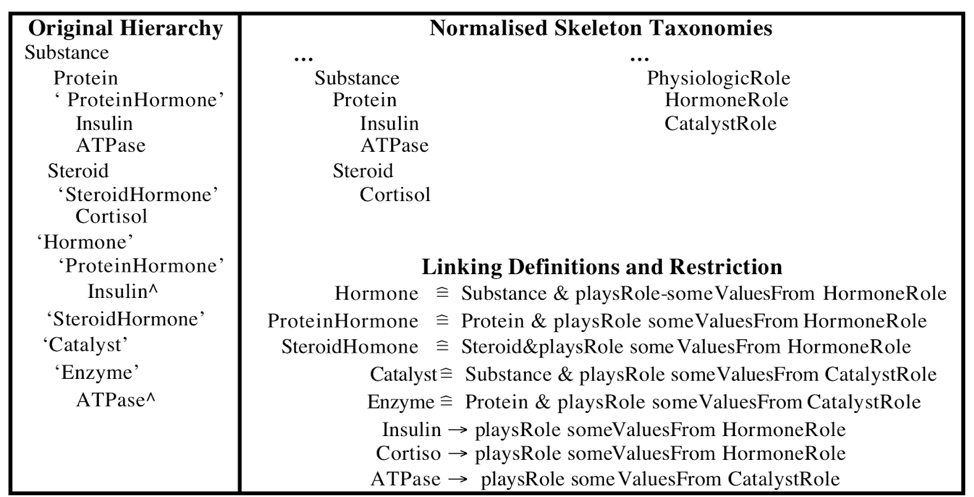

Rector 2003, Figure 1. Original tangled polyhierarchy on the left (multiple parents indicated with “^”). Normalized “primitive skeleton” trees in top left, logical axioms on bottom right. The 3 bars means “equivalent to”, these are logical definitions providing necessary and sufficient conditions. The arrows indicate subClassOf, i.e. necessary conditions. The original hierarchy can be entirely recreated using the skeleton taxonomies, domain axioms, through the use of a reasoner.
Rector calls this approach implementation normalization. The concept of database normalization should be familiar to anyone who has had to create or maintain relational database schemas (one example of how patterns from software and database engineering translate to construction of ontologies; see previous post).
From the paper:
The fundamental goal of implementation normalisation is to achieve explicitness and modularity in the domain ontology in order to support re-use, maintainability and evolution. These goals are only possible if:
- The modules to be re-used can be identified and separated from the whole
- Maintenance can be split amongst authors who can work independently
- Modules can evolve independently and new modules be added with minimal side effects
- The differences between different categories of information are represented explicitly both for human authors’ understanding and for formal machine inference.
Rector describes five features of ontology languages that are needed to support normalized design:
- Primitive concepts described by necessary conditions
- Defined concepts defined by necessary & sufficient conditions
- Properties which relate concepts and can themselves be placed in a subsumption hierarchy.
- Restrictions constructed as quantified role -concept pairs, e.g. (restriction hasLocation someValuesFrom Leg) meaning “located in some leg”.
- Axioms which declare concepts either to be disjoint or to imply other concepts.
Some of the terms may seem unfamiliar due to terminological drift in the ontology world. Here ‘concepts’ are classes or terms, ‘necessary and sufficient conditions’ are sometimes called ‘logical definitions’, or equivalence axioms (represented using ‘intersection_of’ in obo syntax), ‘properties’ are relations (ObjectProperties in OWL), Quantified role-concept pairs are just simple relational class expressions (or simply “relationships” to those coming from OBO-Edit).
The constructs described here are exactly the ones now used in ontologies such as the Gene Ontology and Phenotype Ontologies for construction of logical definitions (see 1, 2, 3). These ontologies have undergone (or are still undergoing) a process of ‘de-tangling’. Many ontologies now employ a prospective normalized development process, where classes are logically defined at the time of creation, and their placement inferred automatically. Examples include uPheno-compliant ontologies such as XPO, ZP, and PLANP. This has the advantage of requiring no retrospective de-tangling, thus saving on wasted effort.
In practice it’s rarely the case that we perfectly adhere to the normalization pattern. In particular, we rarely ‘bottom out’ at pure tree-based ontologies. Usually there is a chain of dependencies from more compositional to less compositional, with the terminal ontologies in the dependency tree being more tree-like. It should also be noted that the practice of normalization and avoidance of assertion of multiple is-a parents has sometimes been mistaken for a principle that multiple parents is bad. This misconception is addressed in a separate post.
It is also considerably easier to do this than when Rector wrote this paper. Protege has seen numerous improvements. One game-changer was the advent of fast reasoners such as Elk which reasons over the EL subset of OWL (or a close enough approximation), which is sufficient to cover the constructs described in the Rector paper, and thus sufficient for basic normalization.
We also have a number of systems for creating normalized ontology classes using templates, where the template corresponds to a design pattern. These include Dead Simple OWL Design Patterns, ROBOT templates, TermGenie, and Tawny-OWL. These allow ontology developers to author logical definitions for classes without directly writing any OWL axioms. An ontologist defines the pattern in advance and the ontology developer can simply fill in ‘slots’.

Example template for GO biochemical process classes. From Hill et al (note some relations may have changed). Using a templating system, a curator needs only select the template (e.g. biosynthetic process) and a value for any template values (e.g. X=alanine, a class from ChEBI).
Where are we now?
The Rector 2003 paper ends with “we believe that if the potential of OWL and related DL based formalisms is to be realised, then such criteria for normalisation need to become well defined and their use routine”. Fast forward sixteen years to 2019, and the bio-ontology world is still engaged in a process of untangling ontologies. Although we have made good progress, some ontologies are still tangled, and many continue to assert hierarchies that could be inferred. Why? There may be a number of reasons for this, including:
It’s tempting to hack

It can be tempting to “hack” together an asserted hierarchy as opposed to constructing an ontology in a modular fashion. This is especially true for ontology developers who have not been trained in techniques like modular design. We see this trade-off in software development all the time: the quick hack that grows to unmaintainable beast.
Retrospectively untangling gets harder the larger the ontology becomes. This is a good example of the technical debt concept from software engineering, outlined in the previous post. Even for well normalized ontologies, a tangled remnant remains, leading to ballooning of technical debt.
We’re still not there with tools
Even where we have the tools, they are not universally used. ChEBI is one of the most widely used bio-ontologies, but it currently lacks logical definitions. This is in part because it is developed as a traditional database resource rather than an ontology. Curators used a specialized database interface that is optimized for things such as chemical structures, but lacks modern ontology engineering features such as authoring OWL axioms or integrating reasoning with curation.
Untangling is hard!

Untangling can be really hard. Sometimes the untangling involves hierarchies that have been conceptually baked in for centuries, along with apparent contradictions. For example, the classification of the second cranial nerve as a nerve, being part of the central nervous system, and the classification of nerves as part of the peripheral nervous system (see the example in this post). Trying to tease apart different nuances into a well-behaved normalized ontologies can be hard.
It should be noted that of course not everything in biology is amenable to this kind of normalization. A common mistake is over-stating logical definitions (there will be a future post on this). Still, there are no shortage of cases of named concepts in the life sciences represented in ontologies that are trivially compositional and amenable to normalization.
Sociotechnological issues confound the problem
In principle the Rector criteria that “maintenance can be split amongst authors who can work independently” is a good one, but can lead to sociotechnological issues. For example, it is often the case that the larger domain ontologies that are the subject of untangling receive more support and funding than the primitive skeleton ontologies. This is not surprising as the kinds of concepts required for curators to annotate biological data will often be more compositional, and this closer to the ‘tip’ of an ontology dependency tree. Domain ontology developers accustomed to moving fast and needing to satisfy term-hungry curators will get frustrated if their requested changes in dependent module ontologies are not acted on quickly, necessitating “patches” in the domain ontology.
Another important point is that the people developing the domain ontology are often different from the people developing the dependent ontology, and may have valid differences in perspective. Even among willing and well-funded groups, this can take considerable effort and expertise to work through.
For me the paradigmatic example of this was the effort required to align perspectives between GO and ChEBI such that ChEBI could be used as a ‘skeleton’ ontology in the Rector approach. This is described in Dovetailing biology and chemistry: integrating the Gene Ontology with the ChEBI chemical ontology (Hill et al, 2013).
For example, a nucleotide that contains a nucleobase, a sugar and at least one phosphate group would be described as a carbohydrate by a carbohydrate biochemist, who is primarily concerned with the reactivity of the carbohydrate moiety of the molecule, whereas general organic chemists might classify it as a phosphoric ester.
Both of the above classifications are correct chemically, but they can lead to incorrect inferences when extrapolated to the process hierarchy in the GO. Consider ‘nucleotide metabolic process’ (GO:0009117), ‘carbohydrate metabolic process’ (GO:0005975) and ‘phosphate metabolic process’ (GO:0006796) in the GO. If ‘nucleotide metabolic process’ (GO:0009117) were classified as both is _a‘carbohydrate metabolic process’ (GO:0005975) and is _a ‘phosphate metabolic process’ (GO:0006796) to parallel the structural hierarchy in ChEBI, then the process that results in the addition of a phosphate group to a nucleotide diphosphate would be misleadingly classified as is _a ‘carbohydrate metabolic process’ (GO:0005975). This is misleading because, since the carbohydrate portion of the nucleotide is not being metabolized, biologists would not typically consider this to be a carbohydrate metabolic process.
This situation was resolved by decreasing is-a overloading in ChEBI by the addition of the functional-parent relationship. But the process of understanding these nuanced differences and resolving them to everyone’s satisfaction can take considerable time and resources. Again, it is harder and more expensive to do this retrospectively, it’s always better to prospectively normalize.
Thankfully this was all resolved, and GO is able to leverage CHEBI as a skeleton ontology in the automatic classification of metabolic processes, as illustrated here:
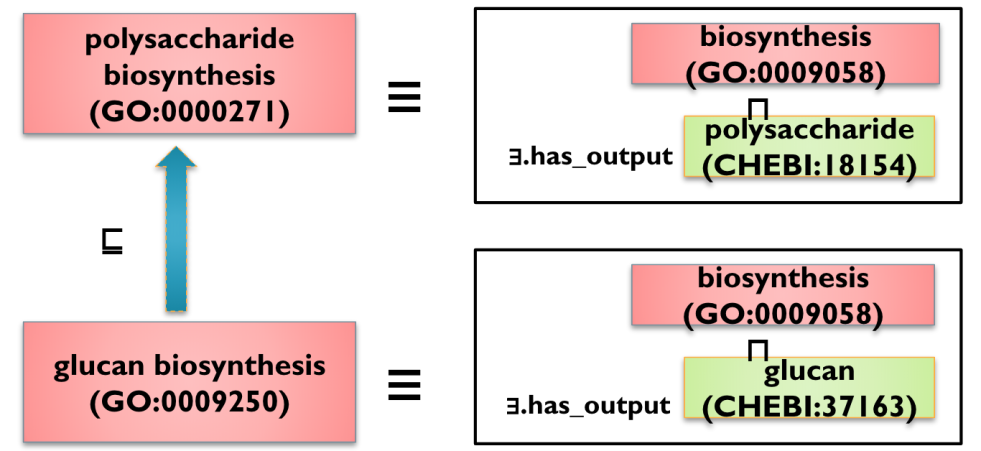
This is one of the motivations for the creation and ongoing operation of the Open Bio-Ontologies Foundry (OBO). One of the motivating factors in the creation of OBO was the recognized need for modular ontology construction and the coordination of the different modules. For me, one of the inspirations for OBO was my involvement in BioPerl development (BioPerl was one of a family of bioinformatics libraries, including BioPython and BioJava). At that time, there was a growing recognition that bioinformatics software was facing a crisis due to unmaintainable code (everyone from that time remembers the “quick perl script” that became enshrined as monolithic infrastructure). The ‘Bio*’ projects attempted to follow best software engineering practice and to break complex code into maintainable modules. Often those modules would be developed by distinct people, but the BioPerl maintainers ensured that these worked together as a cohesive whole.
Summary
- All ontology developers should familiarize themselves with the Rector 2003 paper
- The approach described is particularly useful for larger ontologies that have a high number of compositional classes to manage. Note the vast majority of ontologies in OBO have branches that are to some degree explicitly or implicitly compositional.
- The sooner you adopt this the better – retrospective normalization is harder than prospective.
- Although the approach described is independent of particular technology choice, the adoption of explicit design patterns and a system such as DOSDP-tools, ROBOT templates, or Tawny-OWL is recommended.
- Sometimes the normalization approach involves identifying modules within your own ontology. Sometimes it requires use of an external ontology. This can pose challenges, but rather than give up without trying, use the OBO approach. File tickets on the external ontology tracker, communicate your requirements publicly (it is likely others have the same requirements).
6 thoughts on “OntoTip: Learn the Rector Normalization technique”